Morphological characterization of CCMP470
We characterized the ultrastructure of CCMP470 using light and electron microscopy (LM and EM, respectively). Cells were cylindrical or somewhat barrel-shaped, and were attached to one another by their valve faces to form a chain colony (Fig. 1A,B). A single plastid was recognizable in phase contrast microscopy images (Fig. 1B). The frustules were weakly silicified, as was evident from the observation that critical-point-dried specimens showed a wrinkled surface, particularly in the girdle region (Fig. 1C). Under LM, the width of the lid/bottom of the barrel-shaped cell, which likely represented the diameter of the circular valve, was 2.0 ± 0.7 (1.2–3.4) μm (n = 15). Acid cleaned materials were air-dried, resulting in the collapse of the frustule’s three dimensional structure (Fig. 1D–G), which further confirmed their weak silicification. Girdle bands were numerous (Fig. 1C,D). Although the band surface appeared to be plain in critical-point-dried material observed under SEM, pores were evident in acid-cleaned material observed with TEM (Fig. 1D,F,G). Each band comprised a primary rib running along its long axis (e.g., Figure 1D, arrowhead), from which secondary ribs further extended transversely in both advalvar and abvalvar directions (Fig. 1D, double arrowhead). The secondary ribs were regularly spaced at their bases, resulting in a somewhat regular areolation pattern, but were not fused at the margins (Fig. 1D). The valve had a circular to oblong outline. A sternum ran roughly across its long axis, occasionally showing undulation and/or bifurcation and fusion, which ended up with the formation of an annular structure mostly close to the centre of the valve (Fig. 1E,F, arrow). We interpret this annular structure as a highly reduced labiate process, based on its more or less central position, as well as the fact that its phylogenetic relatives also possess a reduced labiate process in a similar position (see below). We observed valves with more than one annulus along the sternum (Fig. 1G, arrows), although this type of valve can be considered as exceptional, due to its low frequency in the sample. Virgae extended perpendicularly from the sternum, and vimines cross-linked the virgae to form ± square areolae.
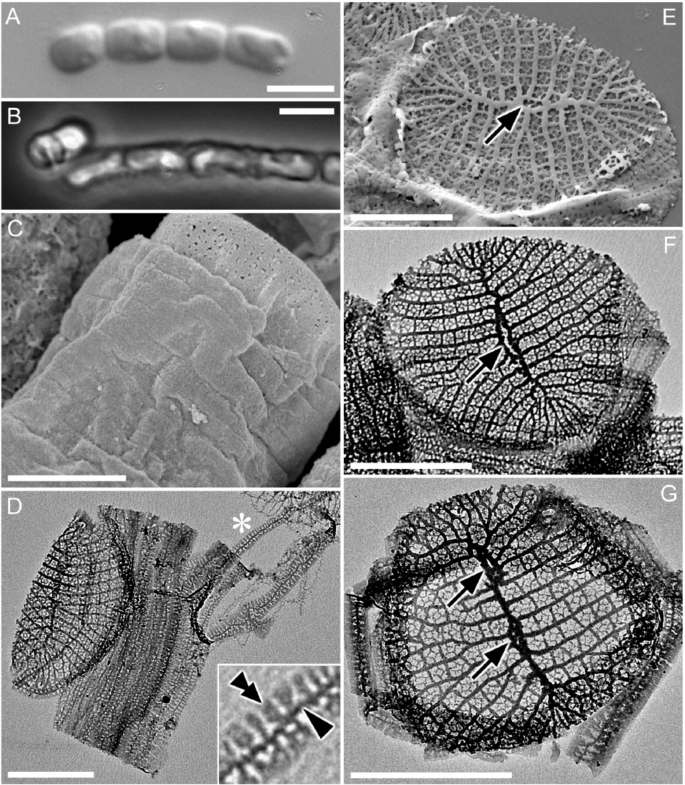

Morphology of CCMP470 under LM (A,B), SEM (C,E) and TEM (D,F,G). (A) Chain colony under bright field, and (B) under phase contrast. (C) Frustule in oblique view, showing numerous girdle bands. Notice the slightly wrinkled appearance of the bands even after preparation by critical-point drying, indicating weak silicification of frustule. (D) Collapsed theca with >10 girdle bands. Each band has a primary rib from which secondary ribs extend perpendicularly. The inset is an enlargement of the part marked by an asterisk, showing the primary and secondary ribs of a band (arrowhead and double arrowhead, respectively). (E–G) Circular valve with a distinct sternum. Virgae extend perpendicularly from the sternum but become radiate towards the periphery of the valve. Occasional bifurcation takes place to keep the stria density even throughout valve face (indicated by arrows), except for valve apices which display a distinct region with slightly finer striae; these most likely represent apical fields responsible for mucilage secretion. The highly reduced labiate process formed along the sternum can be inconspicuous (E), transapically elongated (F), or double (G). Scales = 5 µm (A,B) and 1 µm (C–G).
In LM, the cylindrical cells, attached to one another by their valve faces to form chains, resemble the radial centric genus Leptocylindrus, which explains the former annotation of this strain as Leptocylindrus danicus in the NCMA culture collection. However, the ultrastructure is instead consistent with the staurosiroid clade of araphid pennate diatoms. These are small-celled araphids14,15,16 characterized by the presence of a distinct sternum with parallel striae, features considered as synapomorphies of pennates14,15, and the absence in general of a labiate process on the valve14 (Fig. 1). However, these features are not universally conserved across staurosiroids. The genus Plagiostriata, for example, has an atypical labiate process with an apparently reduced architecture, with no stalk-like structure at its base. Rather, it takes the form of a penetration on the valve face with a distinct marginal area (figs. 14, 15 in15, figs. 330–339 in16). It should be noted in addition that the labiate process in P. goreensis is located at the middle of one side of the sternum, in a position similar to that of the annulus observed in CCMP470 (Fig. 1E,F). There are also some non-staurosiroid araphid diatom genera possessing structurally “modest” labiate processes (i.e., with less elevated architecture and no stalk-like structure at the base) at the centre of the valve, such as Diatoma and Tabellaria17,18. An alternative interpretation for the CCMP470 annulus is that it is an incompletely fused sternum. The pattern centre of the valve in araphid diatoms may perhaps be an elongated annulus that usually gives rise to a sternum by infilling during further valve morphogenesis19. However, because valves with annular structure were dominant throughout the observations using many asynchronous cultures fixed at different times, it is unlikely that they are only present during valve formation. Therefore, the lightly silicified valves of CCMP470 observed in this study were probably mature, leaving the annular part unmasked or incompletely fused. Superficially similar valves can also be seen in chain-forming planktonic centric diatom species such as Subsilicea and Papiliocellulus20.
Draft genome and phylogeny
To better understand the evolutionary placement of CCMP470, we sequenced and assembled a draft genome. The crude CCMP470 genome sequence was separated into two constituent sets of contigs with differing GC contents, with medians of 64% and 51%, respectively (Fig. 2A; Table S1), indicating that it in fact represents a simple metagenome. These contigs were confirmed using MetaBAT21 and last common ancestor (LCA) reconstruction analysis22,23 to correspond to two, different co-sequenced organisms: the diatom host and a bacterial symbiont (Fig. S1Aand Dataset S1). The CCMP470 host genome assembly is 22.99 Mb and consists of 4,992 contigs, containing 8,970 gene models, slightly fewer than the numbers (11,184–34,500) annotated in other diatom genomes (Fig. 2B). BUSCO analysis revealed that the CCMP470 genome was of an equivalent level of completeness (250/303 eukaryotic BUSCOs identified) to the genomes of other diatom species, and was substantially more complete than genome sequences from Thalassiosira oceanica, Pseudo-nitzschia multiseries, and to some extent Synedra acus24,25 (Fig. 2C; S1B). The BUSCO analysis further suggested that there were very few gene duplications in the CCMP470 genome (Fig. 2C).
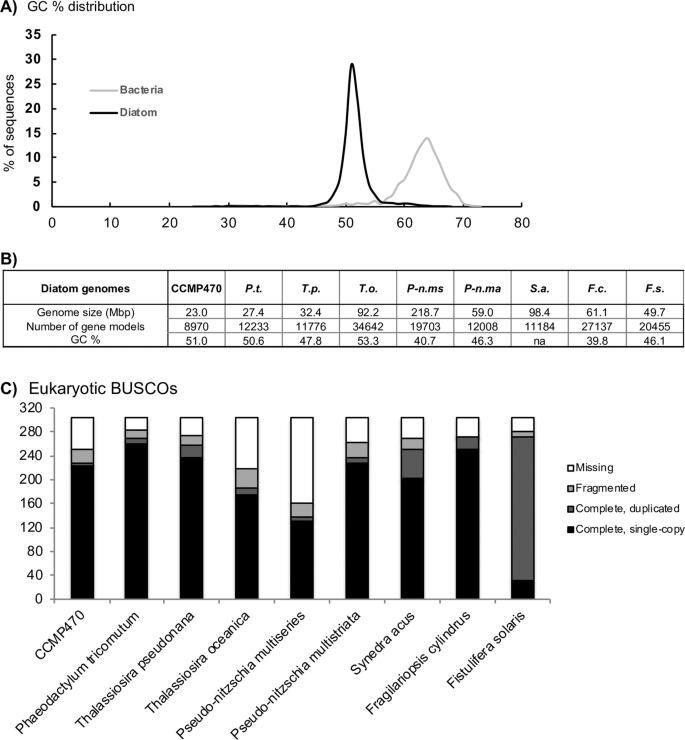
Features of the CCMP470 genome (A) GC content plots of the CCMP470 genome and that of its bacterial symbiont. (B) Comparisons of key features of the CCMP470 genome with those of other diatom species (P.t., Phaeodactylum tricornutum; T.p., Thalassiosira pseudonana; T.o., T. oceanica; P-n.ms, Pseudo-nitzschia multiseries; P-n.ma, Pseudo-nitzschia multistriata; S.a., Synedra acus; F.c., Fragilariopsis cylindrus; F.s., Fistulifera solaris), assembled from24,25,30,31,32,66. “na” indicates the corresponding information was not recorded in the genome release. (C) BUSCO version 3 coverage of different diatom genomes assessed using eukaryotic gene models26; equivalent outputs using plant and protist gene models are shown in Fig. S1B and Table S1.
A concatenated multigene tree was built using a 65 taxa × 16,774 amino acid alignment of 35 eukaryotic BUSCOs identified to have widespread conservation (present in complete or fragmented form in >60% sampled species, average copy number per species <1.5) across all diatoms, following previously established methodology26,27 (Table S2). This topology strongly supported placement of CCMP470 within the araphid pennate diatoms, with a sister-group position to Staurosira robustly supported (posterior probability 1.0) in all three MrBayes trees generated with the concatenated library, and 15 of the 33 single-gene RAxML trees in which it was included (Fig. 3; Table S2).
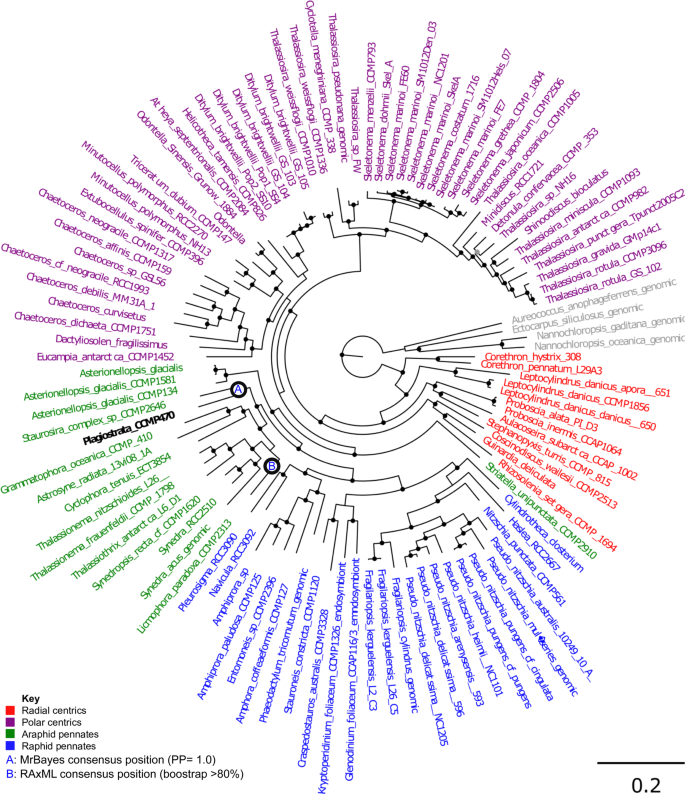
Evolutionary position of CCMP470 based on BUSCO sequences. A. Consensus topology of a 106 taxa × 16,774 aa alignment of 35 BUSCOs conserved in published diatom genome and transcriptome libraries, inferred with MrBayes and RAxML using three substitution matrices (MrBayes: GTR, Jones, WAG; RAxML: GTR, JTT, WAG). The tree topology is rooted on four evolutionary outgroup ochrophyte genomes (in grey). Diatom taxa are shaded by evolutionary origin, and Plagiostriata CCMP470 is shown in black. The topology shown is the consensus MrBayes topology; alternative phylogenetic positions for CCMP470, as inferred using Bayesian and RAxML analysis, are shown with labelled circles.
We also generated a more species-rich phylogenetic tree using a concatenated alignment of SSU, rbcL and psbC produced by Theriot and colleagues7 but with more leptocylindrid and staurosiroid taxa sequenced11,13,14,16, in order to infer the phylogenetic position of CCMP470 with higher resolution (Fig. 4; Table S2). Within the staurosiroid clade, Fragilariforma virescens UTEX FD291 formed a sister-group to all other lineages, with moderate bootstrap support (BS: 84). The rest of the members bifurcated (with BS: 96) into two clades. One with the highest support, comprised Opephora, Staurosira, Hendeyella, Psammotaenia, Pseudostaurosira, Nanofrustulum, Staurosirella and Serratifera, and the other, with no nodal support (bs: <70), contained Synedra, Cratericulifera, Castoridens and Plagiostriata, in which CCMP470 and two Plagiostriata strains (P. baltica SZCZCH1550 and P. goreensis s0388) formed a robust clade, CCMP470 being nested within the Plagiostriata species.
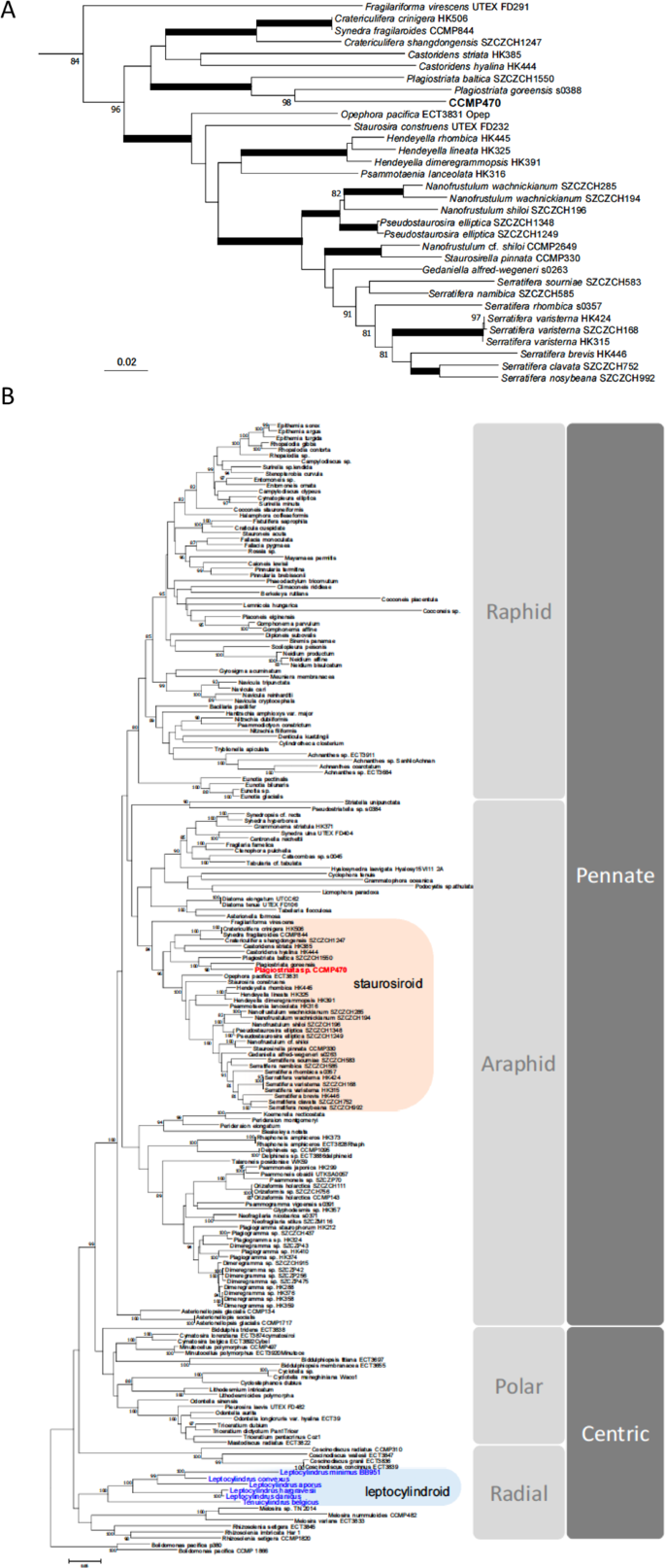
Evolutionary position of CCMP470 based on SSU, rbcL and psbC sequences. (A) Enlarged view of the staurosiroid clade based on a combined dataset of SSU, rbcL and psbC. Nodes with thicker lines indicate 100% bootstrap value. For simplicity only strong nodal support (bootstrap value >80%) is shown. Scale = 0.05 substitutions/site. (B) Complete topology inferred for alignment in A), including non-staurosiroid taxa.
Functional and evolutionary characterization
We wished to determine what functions are encoded in the CCMP470 genome. GO category annotation was generated using protein families (Pfams) found by CLADE28 and refined by GO consortium PFAM2GO29. We found the genome to be enriched in genes encoding functions associated with organelle biogenesis (Golgi apparatus, ER membrane, chloroplast, nuclear chromosome; Fig. S2A). We also identified 270 Pfams that are encoded within this genome, but not found in the genomes of the diatoms Phaeodactylum tricornutum, Thalassiosira pseudonana, T. oceanica, or Fragilariopsis cylindrus (Fig. 5A)24,30,31,32. These include two tetratricopeptide domains (PF07926, PF13429) and a methanol dehydrogenase (PF13360), each found in >5 copies in the CCMP470 genome, but absent from all four remaining diatoms (Fig. S2B). Conversely, we identified 358 Pfams found in all four remaining diatom genomes, but absent from CCMP470, including a pentatricopeptide domain (PF01535) and zinc-binding alcohol dehydrogenase (PF00107), both present in >5 copies in all other diatoms (Fig. S2B).
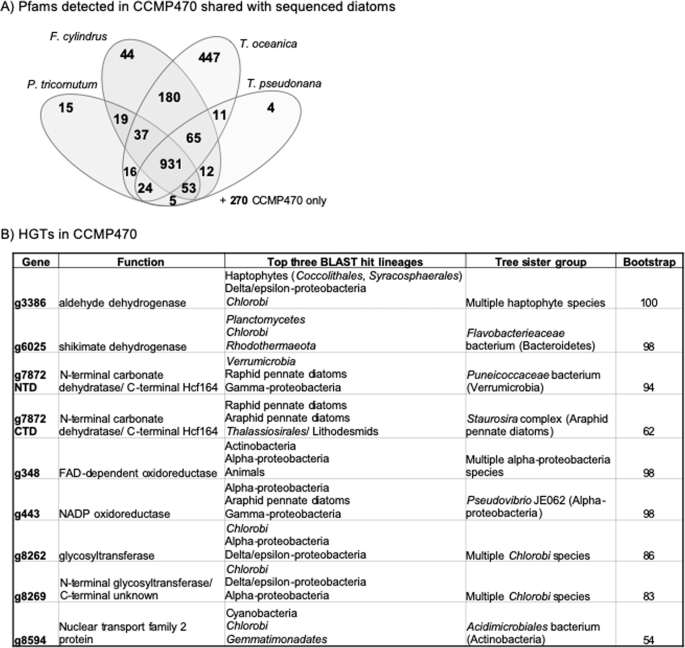

Functional and evolutionary profiling of the CCMP470 genome. (A) Combinations of Pfams found in other sequenced diatom genomes from CCMP470 host genome contigs. 270 Pfams were found uniquely in CCMP470 (shown outside the Venn diagram). GO enrichment data, and specific Pfam domains associated with CCMP470 and not with other diatoms, are shown in Fig. S2B. Individual Pfams for each gene in the CCMP470 genome are provided in Table S1, in the sheet labelled “function”. Tabulated totals of each Pfam for CCMP470, and for each other reference diatom genome, are provided in Table S1 in the sheet labelled “PFAMs”. (B) Evolutionary origins of eight genes verified through both BLAST rank and single-gene tree approaches to have been acquired via horizontal gene transfer into the CCMP470 genome. Exemplar trees and structural diagrams for one protein (g7872) inferred to be a chimera of bacterial and diatom origin are shown in Fig. S4.
Next, we investigated whether any genes in the CCMP470 genome have specific evolutionary ancestries that separate them from other diatom lineages. The evolutionary origin of each gene in the CCMP470 genome was investigated using a reciprocal BLAST best hit (RbH) search, integrating published genomic and transcriptomic data from across the tree of life, and divided into 144 sub-categories based on recently published taxonomies5,23,27 (Table S3). Consistent with the phylogenetic placement, the single diatom taxonomic category with the most reciprocal BLAST best-hits was the araphid pennate diatoms (7,977 genes with RbH matches), followed by raphid pennate species (7,819 genes), with a smaller number of RbH matches (5,974 genes) obtained for Leptocylindrus (Table S3).
We used the outputs of the RbH analyses to search for genes that uniquely possess orthologues in close relatives, following taxonomic nomenclature established by Dorrell and colleagues in33. Through this, we identified five genes that only produced BLAST hits with expect value below 10−05 in other araphid pennate libraries (principally Staurosira complex; Fig. S3A), and thus presumably originated within this lineage. Although none of these genes contained known protein domains, alignments with their homologues revealed large numbers of identical residues, indicating probable functional conservation (Fig. S3B).
We additionally used a previously published phylogenomic pipeline, based on ranking the reciprocal BLAST top hits from each of the 144 sub-categories through BLAST search, followed by single-gene trees, to identify evidence of horizontal gene transfer into a recent ancestor of CCMP470 (Table S4)34. The majority of the genes identified were either most closely related to other diatom sequences, or could not retrieve clear non-diatom origins through the BLAST top hit analysis; or were found to be too divergent to align with their respective BLAST top hits to allow phylogenetic inference. However, we could confidently identify eight candidate HGT events (resolving with a sister group corresponding to the BLAST top hit taxon, with RAxML bootstrap support >50%; Fig. 5B). Seven of the HGT genes are of apparent prokaryotic (chlorobial, alphaproteobacterial, and verrucomicrobial) origin; and the remaining HGT fell in an unresolved position within the haptophytes (Fig. 5B). Seven of the HGT genes occur on contigs containing at least one sequence of verified diatom origin, indicating that they are not contaminants within the diatom sequence fraction (Table S4). The remaining HGT gene (g7872 C) is an apparent chimeric or “S-gene”34, consisting of an N-terminal carbonate dehydrogenase domain of apparent verrucomicrobial origin; and a C-terminal Hcf164 domain of diatom origin (Fig. S4).
Origin and function of the CCMP470 symbiont
Next, we considered the genome of the CCMP470 bacterial symbiont, which was 3.48 Mb in size and consisted of 3,158 genes (Table 1A; Table S1 and Dataset S1). The majority of the bacterial contigs were identified using LCA annotations as resolving within the Rhodobacteraceae, an alpha-proteobacterial group frequently identified as algal symbionts and associated with algal blooms35,36,37,38,39. We used a set of 27 genes from a list of 40 previously identified as single-copy phylogenetic markers for alpha-proteobacteria40, that could be detected on the symbiont contigs (Table S2), to further identify sequences of probable alpha-proteobacterial commensal origin in six other published algal genomes and transcriptomes41,42 (Table 1B).
We also performed reciprocal BLAST best hit and BLAST rank analyses of the symbiont genome, as we had done for the diatom, to identify genes that might have arisen in the bacteria via recent horizontal gene transfer (Fig. S5A). We identified two genes likely to represent genuine HGT events by single-gene trees, arising, respectively, from gamma-proteobacterial and verrucomicrobial donors, the first of unknown function and the second belonging to the abortive infection phage resistance (AIPR) protein family (Table S4). We noted a small number of genes that, based on BLAST rank analyses, had closest evolutionary relatives amongst diatoms and other stramenopile lineages (Fig. S5B). Single-gene phylogenies indicate that these genes frequently have alpha-proteobacterial second sister groups, reinforcing the presence of bacterial symbionts within these cultures (Table S4).
In silico analysis of metabolic interactions between CCMP470 and its bacterial symbiont
Since previous studies have shown that CCMP470 can form symbiotic interactions for nutrient exchange with Rhodobacteraceae, e.g., supplying fixed organic carbon to the species Planktotalea frisia43, we explored the potential of metabolite exchange between CCMP470 and its co-sequenced symbiont in silico. We searched for possible metabolic interactions between the two species using ModelSEED44. This identified 1,201 reactions based on 768 annotated genes for the bacterium, and 1,084 reactions from 502 genes for the diatom (Table S5 and Dataset S2). We noted that the bacterial commensal encodes the complete pathway for the synthesis of bacteriochlorophyll, from geranylgeraniol and chlorophyllide precursors, suggesting that it is likely to be photo-heterotrophic. Previous studies of algal-bacterial symbioses have experimentally evidenced B-vitamins as mediators of symbiotic interactions, e.g., in the model laboratory system involving Lobomonas rostrata and Mesorhizobium loti45, and the marine species Ostreococcus tauri and Dinoroseobacter shibae46. We found no evidence of B-vitamin auxotrophies in either CCMP470 or its symbiont, as both organisms encode complete pathways for thiamin, niacin, folate and biotin biosynthesis. CCMP470 is unlikely to require cobalamin (vitamin B12) obligately, as it encodes a putative cobalamin-independent methionine synthase (METE, 5 methyl-tetra-hydropteroyl-triglutamate-homocysteine methyltransferase, g2483.t1)47; and indeed we could not identify a complete cobalamin biosynthesis pathway in the symbiont. We did note that the symbiont retains a gene encoding BluB, which adds the DMB moiety to pseudocobalamin, forming cobalamin, which would imply that the species is able to use B12 if it can acquire intermediates from its environment48. The ModelSEED analysis predicted that the symbiont is able to process dehydroepiandrosterone sulfate (a sulfate ester), and a range of sulfite and sulfate compounds; and previous studies have shown that certain Rhodobacteraceae can metabolize organosulfur compounds produced by algal commensals49. However, the poor annotations in CCMP470 of osmolyte producing pathways of diatoms did not allow us to evaluate which organosulfur compounds this diatom is able to synthesize, in order to be able to infer metabolic exchanges.
Source: Ecology - nature.com


