We conducted this study from December 4, 2012 through March 12, 2013 with the cooperation of 35 feedlots. Feedlots were located within 4 regions; Eastern Colorado (n = 8), Kansas (n = 8), Texas panhandle (n = 8), Southern Iowa/Northern Missouri (n = 11). Feedlots experiencing bird damage (large foraging flocks of birds) were identified with the help of local cattlemen’s associations. Bird damaged feedlots were randomly selected from a pool of commercial facilities, within each region, that had reported severe bird damage the previous year. Comparable reference feedlots were selected within each geographical region. A total of 18 treated and 17 reference facilities were included in the analysis. All participating facilities group housed animals in pens and produced feeder cattle as their primary commodity. Dairies, calving, or non-cattle livestock facilities were not included in the study.
This experimental protocol was approved by the USDA/APHIS/Wildlife Services, National Wildlife Research Center prior to data collection (Study Director James C Carlson; NWRC Protocol number QA-1945). Starling control operations were conducted by biologists from the United States Department of Agriculture/APHIS/Wildlife Services. Starling control was conducted following agency policy as stated in USDA/APHIS/WS Directive 2.505. All methods were carried out in accordance with relevant guidelines and regulations.
Wildlife Services biologists initially established starling feeding sites within the 18 treatment feedlots using a bait preferentially selected by European starlings. Once starlings were observed to be consistently feeding on pre-bait, biologists used a 2% solution of DRC-1339 (3-chloro-p-toluidine hydrochloride) to reduce the number of depredating starlings. Technical DRC-1339 powder was mixed with water to create a 2% solution. Starling feed was soaked in the 2% solution and screen dried. The bait was applied at a concentration of 1:10 treated to untreated starling feed particles. All DRC-1339 applications were implemented consistent with directions “Compound DRC-1339 Concentrate – Feedlots”; EPA registration 56228–10.
Each feedlot was sampled twice, once before and once after starling control operations. During each sampling period we collected European starlings (n = 30) and cattle feces (n = 50) from within feedlots. Within each feedlot up to 10 pens were selected. These same pens were sampled before and after starling control operations. Within each pen we collected a minimum of five cattle fecal samples per visit. If a feedlot had fewer than 10 pens the total number of samples was distributed, as evenly as possible, among the available pens. For example, one facility housed animals in 2 large pens. Within this feedlot we collected 25 fecal samples per pen per visit. Within some feedlots fewer than 30 starlings were collected if birds could not be found.
Collection of cattle fecal samples followed methods that have been described previously12. Cattle fecal samples were collected from the floor of animal pens and only freshly voided fecal pats were sampled. In other words, the sample was collected from a fecal pat only after an animal was observed defecating. This procedure allowed us to standardize environmental exposure time among fecal samples and estimate herd prevalence of ciprofloxacin-resistant E. coli without confining animals for collection of rectal samples. Ten gram samples were scraped from the top of the fecal pat with disposable plastic spoons and stored in sterile Whirl-Paks (Nasco, Fort Atkinson, WI). We only collected fecal samples if we could reasonably determine, by visual inspection, that the sample was fresh and free of external environmental contaminants. All fecal samples were stored in coolers until they were shipped to the laboratory. Estimates of number of birds in animal pens were collected at the same time as fecal sample collection.
Number of birds on feedlots were estimated using counts of bird numbers on each pen’s floor, feed bunkers, water troughs and feed lanes in front of the sampled pen. Estimates from these four locations were summed to calculate the total number of birds within pens. We averaged the total number of birds within pens among all the sampled pens within a feedlot. This mean bird estimate was multiplied by the total number of pens within the facility to produce a facility level bird estimate.
All starlings were collected with shotguns and no birds were collected off feedlots. All starlings were collected from within the animal pens and pen lanes. Starling samples were collected opportunistically and only done when it was safe to discharge firearms in feedlots. All specimens were individually bagged in sterile Whirl-Paks and stored in coolers until shipping.
Within each facility, diagnostic samples (starlings and cattle fecal samples) were collected on the same day and samples were shipped priority overnight to testing laboratories in Iowa and Colorado. All samples were shipped, in insulated boxes packed with Ice-Brix (Polar Tech Industries, Genoa, IL), to laboratories for isolation of ciprofloxacin-resistant E. coli. Only samples received by the laboratories within 24 hours of the date of collection were screened for ciprofloxacin-resistant E. coli. European starlings were shipped to the United States Department of Agriculture, National Wildlife Research Center (NWRC) in Fort Collins, Colorado, USA. Cattle fecal samples were shipped to Ohio State University, Food Animal Health Research Program in Wooster, OH, USA.
All European starling dissections occurred at the NWRC and were conducted using published methods13. Starling lower gastrointestinal tracts (GI, duodenum to the cloaca) were removed and placed in a sterile Whirl-Paks. To reduce risk of cross-contamination, we saturated the starling carcass, scissors, scalpels, and lab stations with 70% ethanol before removal of each starling GI tract. Lab mats and gloves were replaced after processing each starling. The starling GI samples were macerated for 120 sec at 200 rpm using a Stomacher 80 Biomaster (Seward Laboratory Systems, Bohemia, NY) paddle blender. Fecal material from the macerated starling GI tracts was squeezed by hand to one corner of the bag and an aliquot was extracted using sterile cotton swabs, making sure to completely saturate the tip of the swab. Starling fecal material, on the saturated cotton tipped swab, was then used for direct plating onto selective media.
Starling GI and cattle fecal samples were inoculated onto MacConkey agar (HiMedia, Mumbai, India) supplemented with 1 µg/ml ciprofloxacin (Sigma-Aldrich, St. Louis, MO) using sterile cotton-tipped applicators and incubated at 37 °C for 18–24 hr. Colonies displaying typical E. coli morphology were transferred to 10 ml of tryptic soy broth (TSB) and incubated overnight at 37 °C for 18–24 hours. Species confirmation for starling GI samples was achieved using the API 20E system (bioMérieux, Marcy-l’Étoile, France). E. coli susceptibility to ciprofloxacin was confirmed using the disk diffusion method following Clinical and Laboratory Standards Institute protocols and guidelines (CLSI, 2008). Species confirmation for cattle fecal samples was conducted using lactose and indole tests. All lactose and indole positive isolates were cultured on MacConkey agar supplemented with 2 µg/ml ciprofloxacin. Colonies growing on the agar were isolated and tested for both gyrA and parC chromosomal mutations by PCR using previously reported primers14. PCR products were bi-directionally Sanger sequenced and the resulting data were aligned to the corresponding reference gene sequences available in NCBI Genbank (gyrA gene ID: 946614; parC gene ID: 947499). The gyrA and parC sequences were screened for combinations of chromosomal mutations expected to confer fluoroquinolone resistance15,16 and if they were detected the E. coli isolate was classified as ciprofloxacin-resistant.
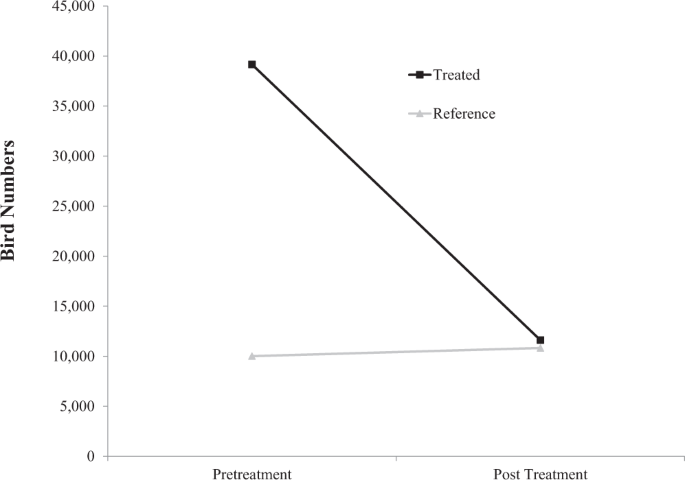
We tested efficacy of DRC-1339 as a control tool for invasive birds on feedlots using a Poisson model of count data in PROC GLIMMIX in SAS version 9.2 (SAS Institute, Cary, NC). The response variable was the estimated number of birds on feedlots. Fixed effects included treatment status (starling controlled feedlot/reference feedlot), sampling period (before/after starling control) and the interaction between treatment status and sampling period. Feedlots nested within treatment status were included as a random effect.
Separate mixed effects logistic regression models were created to test the association between total bird number and ciprofloxacin-resistant E. coli fecal shedding by cattle and to test the efficacy of starling control as a pre-harvest intervention strategy to reduce ciprofloxacin-resistant E. coli fecal shedding by cattle. Models were constructed using PROC GLIMMIX in SAS version 9.2. Both models, were fitted using a binomial distribution and the response variable was the number of positive ciprofloxacin-resistant E. coli samples divided by the total number of samples collected per pen. Model parameters were estimated using the maximum likelihood method and degrees of freedom were estimated using the between within option. Within both models, feedlots nested within treatment status, pens nested within feedlots, and the sampling period by feedlot interaction were all included as random effects.
To test for an association between total bird numbers and cattle fecal shedding of ciprofloxacin-resistant E. coli, we included region and the estimated number of birds on feedlots as fixed effects. To test the efficacy of starling control as a pre-harvest intervention strategy to reduce cattle fecal shedding of ciprofloxacin-resistant E. coli, we included region, treatment status (starling controlled feedlot/reference feedlot), sampling period (before/after starling control operations), and the interaction between treatment status and sampling period as fixed effects.
Additional explanatory variables for ciprofloxacin-resistant E. coli in feedlots were assessed in univariable analyses using PROC GLIMMIX in SAS version 9.2. The model was fitted using a binomial distribution and the response variable was the number of positive ciprofloxacin-resistant E. coli samples divided by the total number of samples collected per pen. Model parameters were estimated using the maximum likelihood method and degrees of freedom were estimated using the between within option. Feedlots nested within treatment status, pens nested within feedlots, and the sampling period by feedlot interaction were all included as random effects.
The additional explanatory variables were assessed to identify any potential wild bird, facility management, or environmental variables that may potentially be associated with cattle fecal shedding of ciprofloxacin-resistant E. coli in feedlots. The explanatory variables assessed in the analyses were selected because they have been identified as or suspected of contributing to bacterial contamination in feedlots13,17,18,19,20. The variables assessed in these analyses occurred at two spatial scales (feedlots and pens within feedlots). The variables include birds (birds in feed bunkers, birds on water troughs, total number of birds in pens), cattle stocking (herd size, number of cattle within pen), environmental factors (temperature, time, and sampling period), and feedlot management factors (water troughs split pens, recycled water used in water troughs, cattle days in pen, cattle days on finishing ration, entry weight, exit weight and weight gained by cattle).
Most variables assessed within the univariable analyses are intuitively obvious, but some variables may need additional clarification. For example, weight gain was calculated by subtracting the pen averaged entry weight from the pen averaged exit weight data. Water troughs accessed by multiple pens identifies split-pen watering troughs that allow cattle from adjoining pens to drink from the same trough. Recycled water identifies facilities that recirculate the water provided to cattle within troughs. Total number of birds per pen reflects the sum of the estimates of birds from water troughs, pen floor, feed bunkers and pen lanes for each sampled pen.
A total of 15 additional univariable models were analyzed (m = 15). Because multiple tests were being conducted, we decided to control for false discoveries using the Benjamini Hochberg procedure21. For all univariable analyses the false discovery rate was set at α = 0.05. Models were ranked by p-values from smallest (1) to largest (m). Cutoff values for rejection of null hypotheses were calculated as (rank/m)*α. Reported odd ratios and their 95% confidence intervals were not adjusted for multiple testing.
Source: Ecology - nature.com



