Screening, purification, and identification of deep-sea bacteria possessing potential thiosulfate oxidation capability
During the course of screening of deep-sea bacteria possessing potential thiosulfate oxidation capability, strain 21–3 attracted our attention because it produced an obvious milky substance in the culture supplemented with 40 mM thiosulfate after two days of incubation (Fig. 1a), which is a typical indicator of ZVS formation. The cells of strain 21–3 were subsequently purified three times using the dilution-to-extinction technique at 28 °C. The 16S rRNA gene sequence of strain 21–3 (accession no. MN744319) shared a high similarity of 99.93% with Erythrobacter flavus SW-46T. In addition, strain 21–3 also clustered with E. flavus SW-46T according to phylogenetic analysis (Supplementary Fig. S1). The ANI between strain 21–3 and other two strains E. flavus VG1 (accession no. ASM223761) and E. flavus KJ5 (accession no. ASM429625) was 97.56% and 97.40%, respectively, which were higher than the accepted threshold (ANI value of 94%) for same species [55] Thus, strain 21–3 was identified as a member of E. flavus and designated as E. flavus 21–3 in this study.
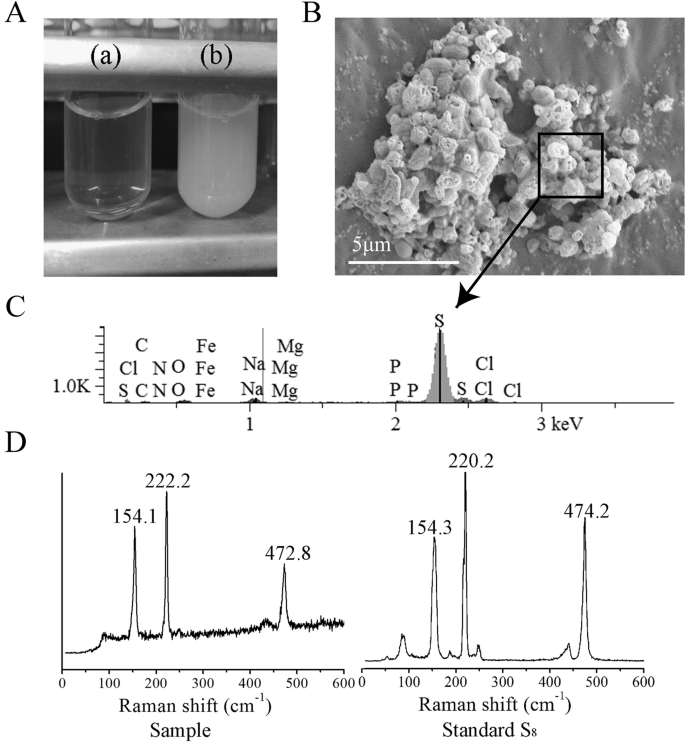
E. flavus 21–3 cultured in diluted 2216E without (a) or with 40 mM thiosulfate (b). b SEM observation of ZVS produced by E. flavus 21–3. c Energy dispersive spectrum analysis of the selected area in b. d Raman spectra of the ZVS produced by E. flavus 21–3 and standard S8.
Identification and characterization of sulfur globules produced by E. flavus 21–3 combining with SEM, EDS, Raman spectrum, and TEM
To gain insight into the milky white substance produced by E. flavus 21–3 when cultured with thiosulfate, the white substance was subjected to SEM and EDS analyses. SEM results showed that bacterial cells were embedded in globular crystals (Fig. 1b), which is similar to the reported extracellular sulfur globules produced by Hydrogenovibrio thermophiles, a chemolithomixotrophic bacterium that uses thiosulfate as an energy source [56]. These crystals were further identified as elemental sulfur by EDS (Fig. 1c). Raman spectrum of the cyclooctasulfur standard was characterized by two strong peaks at 154 and 220 cm−1 corresponding to the bending and stretching modes of the eightfold ring and a third peak at 473 cm−1 [4, 10]. These typical characteristics of S8 were also found in the analysis of the white substance from E. flavus 21–3 (Fig. 1d).
To further characterize the ZVS intermediate formed in E. flavus 21–3, sulfur globules produced by the cells were purified and checked with TEM. TEM results showed that the separated sulfur globules (of size 400 nm) were coated with carbon (Fig. 2a) as previously described [57]. TEM-EDS results of the separated sulfur globules (Fig. 2b) were accordant with those of SEM–EDS (Fig. 1c), confirming that E. flavus 21–3 is capable to convert thiosulfate to ZVS. TEM was further conducted to observe the production of sulfur globules (S8) throughout the growth period of E. flavus 21–3 that was cultured with 40 mM thiosulfate. At the start of the early-exponential phase (12 h of incubation) many small solid globules were produced around the cells (Fig. 2c), while at the mid-exponential phase (cultured for 36 h) only a few large, hollow globules were attached to the cells (Fig. 2d). These distinct morphological characteristics of the sulfur globules observed at different growth stages suggest that the sulfur globule may serve as an intermediate during the oxidation of thiosulfate, and indicated that E. flavus 21–3 might be a good candidate for the study of microbial thiosulfate oxidation.
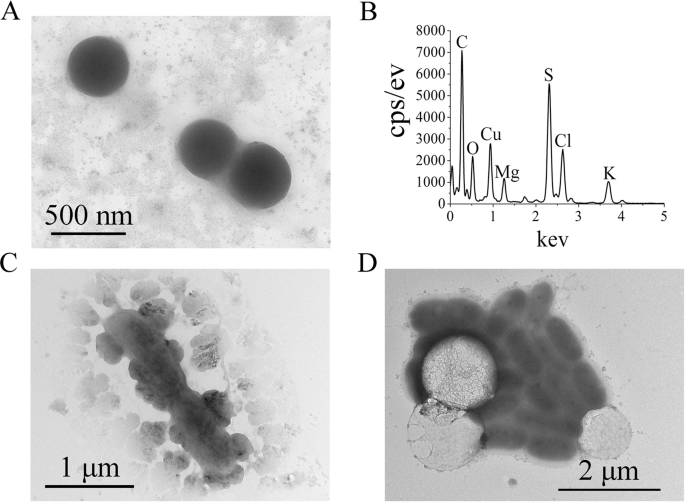
a TEM image of purified ZVS produced by E. flavus 21–3. b Energy dispersive spectrum analysis of the separated ZVS shown in panel A. TEM images of E. flavus 21–3 cells mixed with ZVS when cultured for 12 h (c) or 36 h (d) in medium supplemented with 40 mM thiosulfate.
Genomic evidence of thiosulfate oxidation in E. flavus 21–3
To obtain a genetic basis of the thiosulfate oxidation in E. flavus 21–3, the genome of E. flavus 21–3 was completely sequenced using PacBio RSII (Novogene Bioinformatics Technology Co. Ltd, Beijing, China), and the general genomic features are presented in Supplementary Table S3. The complete genome sequence and all genome information of E. flavus 21–3 have been deposited to GenBank (accession no. CP032228). When analyzing the E. flavus 21–3 genome sequence, we surprised to find only some genes encoding sulfotransferase and thioredoxin-disulfide reductase present, such as disulfide interchange, sulfite exporter, and proteins involved in the metabolism of assimilatory sulfate reduction (Supplementary Table S4). While most typical genes encoding proteins involved in thiosulfate oxidation (the Sox multienzyme complex, flavin cytochrome c-sulfide dehydrogenase, sulfide: quinone oxidoreductase, sulfite: acceptor oxidoreductase, and reverse-acting dissimilatory sulfite reductase) were absent. Given the fact that thiosulfate oxidation occurs in E. flavus 21–3, we speculated that a novel pathway may exist in E. flavus 21–3.
Proteomic analyses of thiosulfate oxidation in E. flavus 21–3
To better describe the mechanism of thiosulfate oxidation in E. flavus 21–3, a proteomic assessment of E. flavus 21–3 cultured with or without thiosulfate was performed. The results of this assay showed that 346 of 1068 and 508 of 948 proteins were significantly up-regulated when grown in culture containing thiosulfate at 18 and 36 h, respectively (P < 0.05). After carefully analyzing this data, we found that the expression of a gene cluster including genes from D0Y83_01340 to D0Y83_01450 (locus tag) was upregulated invariably at both culture time points (Fig. 3a).
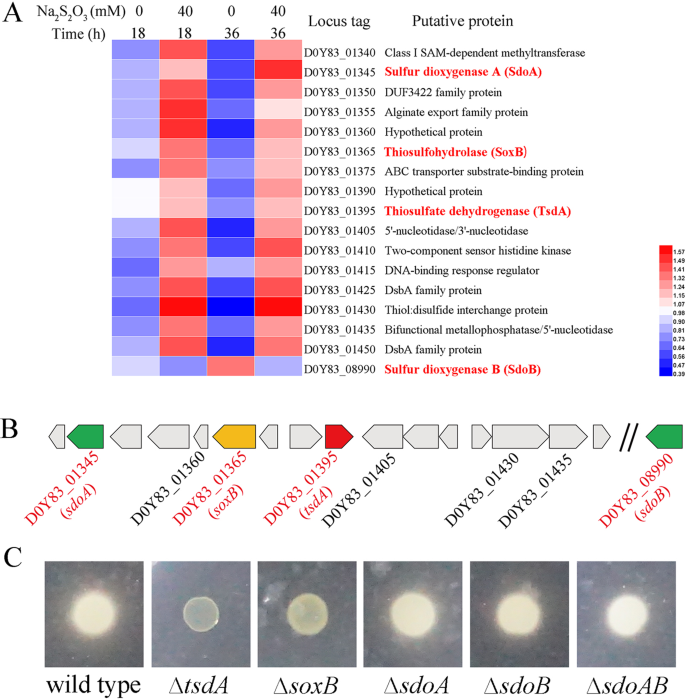

a Heatmap analyses of different expressed proteins in E. flavus 21–3 when cultured in medium supplemented with 0 or 40 mM thiosulfate after 18 h and 36 h, respectively. Expression of D0Y83_08990 (sdoB) is also shown for its high similarity with D0Y83_01345 (sdoA). b Arrangement of the corresponding gene cluster together with D0Y83_08990 (sdoB), possibly involved in thiosulfate oxidation as shown in a. c Genetic determination of genes involved in thiosulfate oxidation as shown in a and b. The capacity of E. flavus 21–3 wide type and corresponding mutants to produce extracellular sulfur globules was assessed using agar plates supplemented with 40 mM thiosulfate. The formation of a white milky substance around the colonies indicates they are capable of thiosulfate oxidation, while the disappearance of the substance around the colonies indicates this capacity has been lost. The genes responsible for thiosulfate oxidation in E. flavus 21–3 (verified by gene disruption) are indicated in red in a and b.
Within this gene cluster, the gene D0Y83_01395 encodes a protein homologous to the c-type cytochrome with C××CH motifs. Since SoxA and TsdA, the typical thiosulfate oxidation proteins, both have the C××CH motif (Supplementary Fig. S2), a phylogenetic analysis was carried out based on the alignment of typical SoxA and TsdA protein sequences. Protein encoded by the gene D0Y83_01395 clustered with TsdA (Supplementary Fig. S3a), which has been shown to play a key role in the formation of tetrathionate from thiosulfate [58, 59]. Thus, the name tsdA was designated to the gene D0Y83_01395, with the corresponding protein named TsdA. Given the importance of TsdA in the course of thiosulfate oxidation, we overexpressed and purified the TsdA of E. flavus 21–3 in E. coli. The apparent molecular mass of purified E. flavus 21–3 TsdA is 36 KD (Supplementary Fig. S3b), larger than the homologous protein in A. vinosum (27 KD) [60]. The UV–Visible electronic absorption spectrum of the purified recombinant protein has an obvious absorption at 410 nm (Supplementary Fig. S3c), a typical indicator of the existence of cysteine-ligated heme in TsdA [58, 59], which confirms that the D0Y83_01395-encoded protein is a homolog of TsdA.
Within the upregulation gene cluster, proteins encoded by genes D0Y83_01365 and D0Y83_01435 were both annotated as “bifunctional metallophosphatase/5′-nucleotidases”, and the protein encoded by gene D0Y83_01405 was annotated as “5′-nucleotidase/3′-nucleotidase”. Further BLASTP results showed that all three proteins possess a conserved domain named “thiosulfohydrolase SoxB”, which is proposed to play a potential role in the cleavage of terminal sulfone groups [61]. The phylogenetic analysis (Supplementary Fig. S4a) and sequence alignment (Supplementary Fig. S4b) of D0Y83_01365, D0Y83_01405, and D0Y83_01435 encoding proteins with other published typical SoxBs further indicate that these three proteins might play SoxB-like function in E. flavus 21–3.
The protein encoded by gene D0Y83_01345 is annotated as “MBL fold metallo-hydrolase” and possesses conserved polysulfide dioxygenase and rhodanase, which are proposed to involve in the oxidation of polysulfide and sulfur [62, 63]. In addition, gene D0Y83_08990 (beyond this cluster) encodes a protein that is highly similar to the protein encoded by gene D0Y83_01345 (Supplementary Fig. S5), also suspected to be involved in the oxidation of polysulfide and sulfur. The arrangement of the gene cluster encoding up-regulated proteins, together with gene D0Y83_08990 is shown in Fig. 3b.
Genetic determination of key factors responsible for thiosulfate oxidation in E. flavus 21–3
Next, we sought to thoroughly investigate the roles of the key genes identified in the proteomic analysis by creating gene knockouts in vivo. After much effort, a genetic operating system was successfully constructed for the first time in the genus Erythrobacter. Notably, deletion of tsdA (D0Y83_01395) and D0Y83_01365 almost completely disrupted the ability of E. flavus 21–3 mutants to form sulfur globules when grown in the presence of thiosulfate (Fig. 3c), which demonstrates that these two genes play key roles in thiosulfate oxidation in E. flavus 21–3. Given the close phylogenetic relationship between D0Y83_01365 encoding protein and other SoxBs (Supplementary Fig. S4a), the gene D0Y83_01365 was designated as soxB with the corresponding protein name SoxB. Though D0Y83_01405 and D0Y83_01435 encoding proteins also possess the “thiosulfohydrolase SoxB” domain, deletion of D0Y83_01405 and D0Y83_01435 did not affect sulfur globule formation (Supplementary Fig. S6).
Deletion of D0Y83_01345 and D0Y83_08990 had no obvious influence on sulfur globule formation (Fig. 3c). However, compared with wild type, the ability of further ZVS oxidation was abolished in the mutant strain ΔD0Y83_08990, which suggests a potential function for D0Y83_08990 encoding protein in ZVS oxidation. Given the high similarity of D0Y83_01345 and D0Y83_08990, we assigned SdoA (sulfur dioxygenase A) and SdoB (sulfur dioxygenase B) to D0Y83_01345 and D0Y83_08990 encoding proteins, respectively. Consistently, double mutant strain ΔsdoAB showed the same phenotype as that of the mutant strains ΔsdoA and ΔsdoB (Fig. 3c). The deletion of other genes, including D0Y83_01360 and D0Y83_01430, did not show any effect on thiosulfate oxidation (Supplementary Fig. S6), suggesting they might be not essential for thiosulfate metabolism in E. flavus 21–3.
Thiosulfate oxidation in E. flavus 21–3 wild type and mutant strains
To reveal the exact function of key genes (tsdA, soxB, sdoA, and sdoB) responsible for thiosulfate oxidation determined by proteomic and genetic methods, the stoichiometry of sulfur intermediates in the course of thiosulfate oxidation in E. flavus 21–3 wild type and mutants was determined. In wild type E. flavus 21–3, the concentration of thiosulfate decreased with the culture time, and reached zero after 48 h of incubation (Fig. 4). Meanwhile, the concentration of sulfate continued to increase throughout the culturing time (~60 h), suggesting that thiosulfate was eventually converted to sulfate. The maximum accumulation of ZVS (S8) in the medium appeared at 24 h and then dropped gradually to the detectable amount at the end of culturing time (Fig. 4), while the concentration of the other important intermediate tetrathionate increased slightly throughout the time course of the culture (Fig. 4). The existence of S8 and tetrathionate in the medium implies that E. flavus 21–3 mediated incomplete oxidation of thiosulfate (Fig. 4). The pH of medium decreased across the culturing time, which strongly indicates the occurrence of sulfone oxidation in the course of thiosulfate oxidation in E. flavus 21–3 [61].
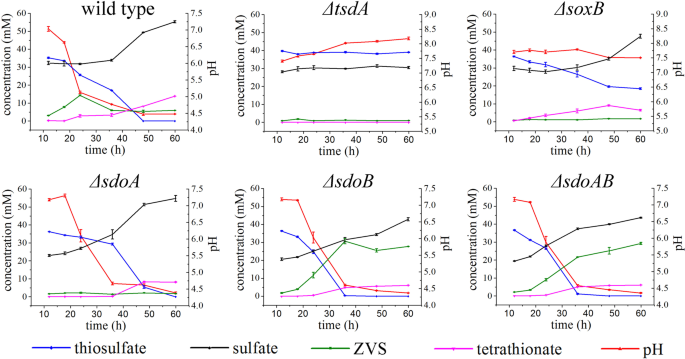
The error bars indicate the standard deviation (S.D.) from two different biological replicates.
Interestingly, the concentration of thiosulfate was constant and tetrathionate was not detected in the ΔtsdA mutant culture (Fig. 4). Given that thiosulfate is generally converted to tetrathionate [12], and the fact that tetrathionate was detected in wild type E. flavus 21–3 but not in the ΔtsdA mutant, it is reasonable to deduce that TsdA mediates the first step of thiosulfate oxidation in this bacterium. It therefore makes sense that neither ZVS nor extra sulfate was detected in the ΔtsdA mutant culture medium. In the mutant strain ΔsoxB, the concentrations of thiosulfate and tetrathionate gradually decreased and increased respectively, while ZVS was not detected in the medium. Meanwhile, the sulfate concentration was lower in the mutant than that in the wild type, indicating the involvement of SoxB in liberating sulfone from tetrathionate (−O3S–S–S–SO3−) during sulfate formation. Taken together, these results suggest that SoxB functions downstream of TsdA but upstream of the key step in ZVS formation.
Mutant strain ΔsdoA consumed 40 mM thiosulfate after 60 h of incubation, without obvious accumulation of ZVS. This observation conflicts with the phenotype detected on agar plates (Fig. 3c). However, we also noticed that more ZVS accumulated in the ΔsdoB mutant than in the wild type, suggesting that SdoB might act as a key factor in ZVS metabolism. Quantitative real-time PCR analysis showed that expression of sdoB was upregulated in the mutant ΔsdoA when the bacterium was cultured with 40 mM thiosulfate (Supplementary Fig. S7), which explained the extremely low concentration of ZVS detected in ΔsdoA. Consistently, the concentrations of thiosulfate, sulfate, tetrathionate, and ZVS in the double mutant strain ΔsdoAB showed similar patterns to those in ΔsdoB, confirming the prominent function of SdoB in oxidation of ZVS in E. flavus 21–3.
Based on the combination of proteomic, genetic, and stoichiometric data on thiosulfate oxidation in E. flavus 21–3 we propose a novel thiosulfate oxidation pathway (Fig. 5). Briefly, thiosulfate oxidation in E. flavus 21–3 is initially mediated by TsdA, which converts thiosulfate to tetrathionate [58, 59]. Tetrathionate is further hydrolyzed by SoxB to form sulfate and release H+, which decreases the environment pH. The remaining sulfur atoms may attach to membrane-bound thiol groups or be added to other polysulfide species [10]. However, polysulfide (polythionate) is unstable in acidic conditions and is therefore partly converted to stable cyclooctasulfur S8 [8, 64, 65], forming the putative sulfur globules. In the presence of SdoA/SdoB, sulfur globules are further oxidized to sulfite and then non-enzymatically/enzymatically converted to sulfate.
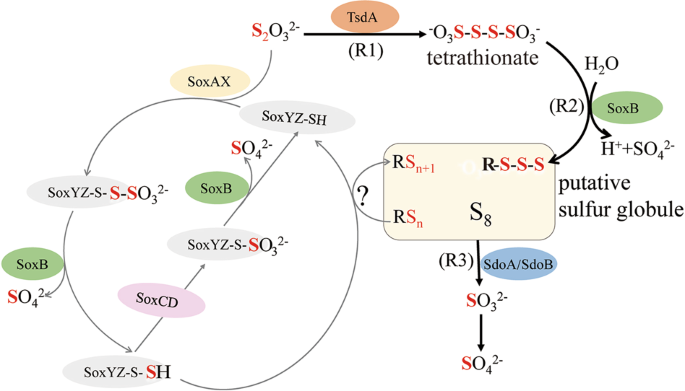
Gray lines indicate the classical Sox pathway and black lines represent the novel pathway determined in E. flavus 21–3. The classical Sox pathway begins with the combination between thiosulfate and SoxYZ with the help of SoxAX, and results in a thiocysteine-S-sulfate residue (SoxYZ-S-SSO32−). Sulfate is liberated by hydrolysis (catalyzed by SoxB) to yield a polysulfide intermediate (SoxYZ-S-S−), which is subsequently oxidized by SoxCD to form a cysteine-S sulfate residue (SoxYZ-S-SO32−). Hydrolysis by SoxB releases sulfate and regenerates the SoxYZ complex (SoxYZ-SH). In the absence of SoxCD, sulfane splitting from the polysulfide intermediate (SoxYZ-S-S−) in an obscure way results in polysulfide and free SoxYZ-SH. The novel pathway found in E. flavus 21–3 begins with the formation of tetrathionate from two molecular thiosulfate, catalyzed by TsdA (Reaction 1: R1). SoxB is responsible for liberating the sulfate from tetrathionate (instead of SoxYZ-S-S-SO3; Reaction 2: R2). The remaining sulfane is converted to polysulfide, and then relatively stable S8. The sulfur globule is finally oxidized by SdoA/SdoB, with sulfite as the intermediate product (Reaction 3: R3). Sulfite is non-enzymatically/enzymatically converted to sulfate. SoxAX sulfur oxidation c-type cytochrome, SoxB thiosulfohydrolase, SoxYZ thiosulfate oxidation carrier protein, SoxCD sulfur oxidation protein. TsdA thiosulfate dehydrogenase, SdoA/SdoB sulfur dioxygenase A/sulfur dioxygenase B.
Wide distribution of the novel thiosulfate oxidation pathway determined in E. flavus 21–3
To explore homologs of TsdA, SoxB, and SdoA/SdoB in other Erythrobacter species, we searched all assembled genomes of Erythrobacter in the NCBI database. Protein sequences PHR02631.1, HAG37069.1, and HAD16915.1 in the Erythrobacter genomes of NORP101, UBA9459, and UBA9044, respectively, showed 99.9% identity with TsdA, and C××CH motifs were also identified in these sequences (Supplementary Fig. S8). Protein sequences HAG37075.1 and HAD16921.1 in the Erythrobacter genomes of UBA9459 and UBA9044 showed 97% identity with SoxB (Supplementary Fig. S9). SdoA/SdoB homologs were found to exist in half of the assembled genomes of Erythrobacter, including in UBA9459 and UBA9044. A phylogenic tree was constructed using SdoA/SdoB sequences from both Erythrobacter and Acidithiobacillus (the typical bacterium possessing ZVS oxidation) (Fig. 6). The results showed that SdoA and SdoB from Erythrobacter were clustered into a separate clade with those from Acidithiobacillus. Given the specific function of SdoA/SdoB in ZVS oxidation, the resulting sequences clustered with SdoA/SdoB were expected to have the same function. Overall, our analysis revealed that homologous sequences of TsdA, SoxB, and SdoA/SdoB were found in assembled Erythrobacter genomes UBA9459 and UBA9044, indicating that this novel thiosulfate oxidation pathway might exist in other Erythrobacter species.
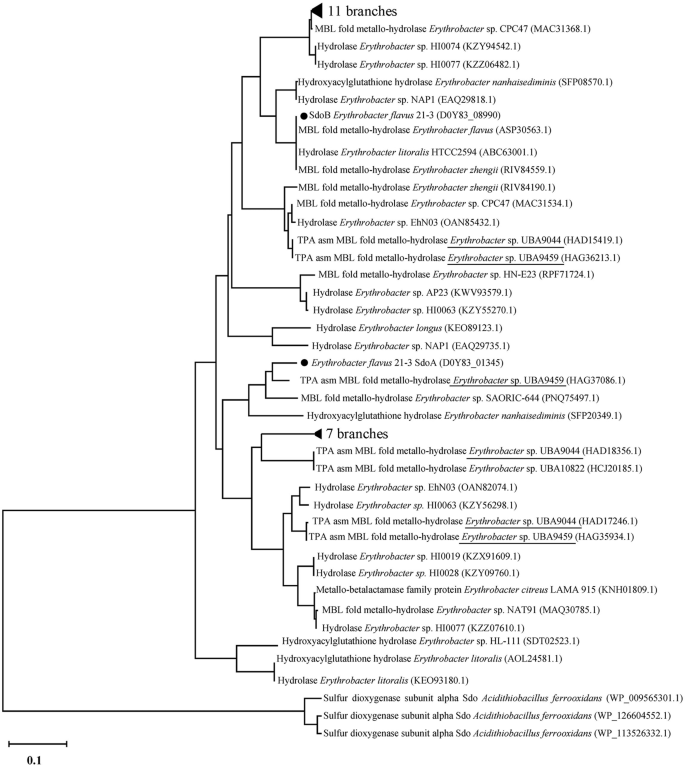
Sequences from Erythrobacter sp. UBA9044 and Erythrobacter sp. UBA9459 are underlined.
Lastly, we checked the bacterial genomes available in IMG database to explore the existence of the novel thiosulfate oxidation pathway in other bacterial species. To ensure the accuracy of this analysis, stringent criteria were applied and only organisms containing both tsdA and soxB homologous genes were considered. The search results showed that the set of genes were mainly identified in Alphaproteobacteria, Betaproteobacteria, Gammaproteobacteria, and Bacteroidetes (Supplementary Table S5). Moreover, the TsdA and SoxB homologs were also searched in the available metagenomes from IMG. The e-value was set as 1e−20 and the sequences with an identity more than 40% were selected. Totally, there were 18 (in total 103) metagenomes containing both TsdA and SoxB homologs (Supplementary Table S6). Notably, the identified bacteria containing both TsdA and SoxB homologs occur naturally in a variety of environments including land soil, acid mine drainage, sea water, and deep-sea sediment. To further clarify the functions of selected TsdA- and SoxB-like proteins, phylogenic trees were constructed using TsdA- and SoxB-like sequences obtained from the identified bacteria (Supplementary Table S5 and Fig. 7). The phylogenetic results showed that these TsdA- or SoxB-like proteins identified in different bacteria clustered with different typical TsdA or SoxB, suggesting that these bacteria might oxidize thiosulfate through the novel pathway. Given the key roles of TsdA and SoxB in this novel pathway and their broad distribution in bacteria, we propose that the novel thiosulfate oxidation pathway identified in E. flavus 21–3 exists in many other microorganisms.
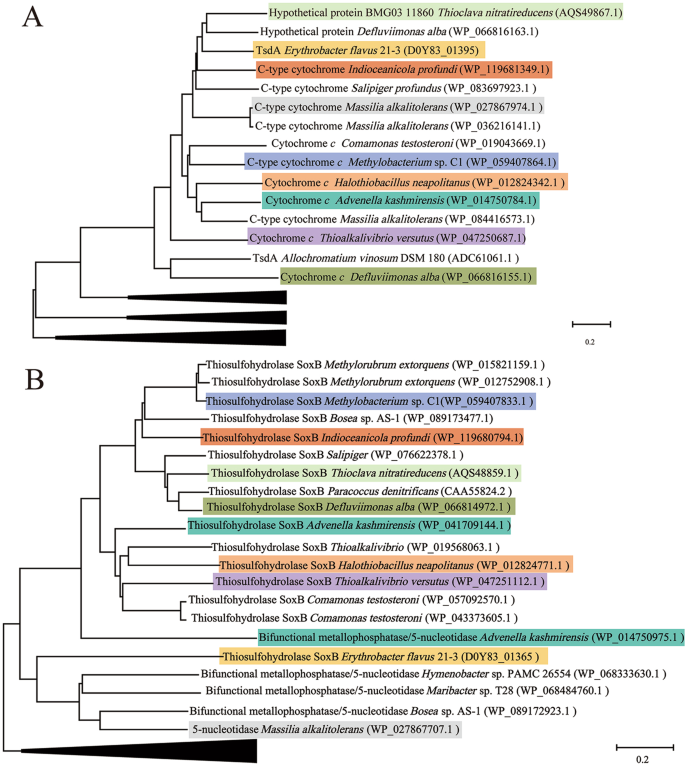
The consensus phylogenetic tree of TsdA (a) and SoxB (b) in E. flavus 21–3 with other homologs obtained from public bacterial genomes (accession numbers are indicated after each species name) constructed by the neighbor-joining method. Branches shown in the tree represent sequences clustered with typical TsdA or SoxB, and each color indicates a specific species in a and b. Related species containing both TsdA and SoxB and their corresponding GenBank accession numbers are listed as follows: Advenella kashmirensis: GCA_000219915.3; Methylobacterium sp. C1: GCA_001854385.1; Indioceanicola profundi: GCA_003568845.1; Thioclava nitratireducens: GCA_000024765.1; Defluviimonas alba: GCA_000024765.1; Halothiobacillus neapolitanus: GCA_000024765.1; Thioalkalivibrio versutus: GCA_001020955.1; Massilia alkalitolerans: GCA_000427785.1, and E. flavus 21–3: GCA_008932225.1.
Source: Ecology - nature.com


