Experimental design
The field experiment was conducted from June 2010 to May 2016 in the Low-Carbon Agricultural Engineering Technology Research Center, Zhuanghang Comprehensive Experimental Station, Shanghai, China (30°53′N, 121°23′E). The site is characterized by a subtropical climate with an average annual temperature of 15.8 °C and rainfall of 1178 mm. Rice (Oryza sativa L.) and wheat (Triticum aestivum L.) were rotated in the experimental field. The soil type was classified as sandy loam with an average SOC concentration of 8.50 g kg−1, total nitrogen (TN) concentration of 0.86 g kg−1, total phosphorus (AP) concentration of 0.52 g kg−1, available nitrogen (AN) concentration of 18.50 mg kg−1, AP concentration of 20.85 mg kg−1, CEC of 15.50 cmol kg−1, and pH of 8.33 in the 0–20 cm soil layer at the beginning of the experiment.
Four treatments were run in triplicate with a randomized complete block design, with each plot area being 60 m2. The crop varieties and management were the same apart from fertilization as follows: CK: non-fertilization; CF: chemical fertilizer application; CS: 3 t ha−1 straw directly returning; CB: 1 t ha−1 straw-derived biochar return. In the wheat growing season, 180, 90, and 204 kg ha−1 of pure N, P, and potassium (K) were applied, respectively; the application amount of pure N, P, and K was 225, 112.5, and 255 kg ha−1, respectively, during the whole rice-growing season. The same amount of total pure N, P, and K was applied in the CF, CS, and CB treatments; the deficiencies were implemented with inorganic fertilizers (Table S1). Inorganic N, P, and K fertilizers were urea, calcium superphosphate, and potassium sulfate, respectively. Pyrolysis was anaerobically performed at approximately 450–600 °C in a vertical charcoal furnace (ECO-5000, Zhejiang, China) to produce biochar. Soil was turned over for all treatments before each crop season planting, during which straw/straw-derived biochar was added into the soil. In detail, after rice/wheat harvest, straw/straw-derived biochar was spread on the soil surface as an amendment and thoroughly mixed with soil at approximately 15–20 cm depth prior to the subsequent wheat/rice crop planting. Therefore, all treatments had the same degree of soil disturbance.
Soil sampling and measurement
Samples were collected from each plot after the wheat harvest on May 8, 2016. They were aseptically collected from 0–20 cm and 20–40 cm depths with a stainless-steel auger (15 mm interior diameter) using the five-point sampling method. The five pools in each plot were combined to provide one composite sample. Samples were placed into sterilized polyethylene sealed bags, stored at low temperature, and then immediately brought back to the laboratory. The fresh soil was gently peeled along natural planes of weakness; visible stones, roots, and other residues were removed. Approximately 100 g of each fresh soil sample was fractionated to different aggregate sizes (as described below) and subsequently frozen at −80 °C for bacterial community analysis.
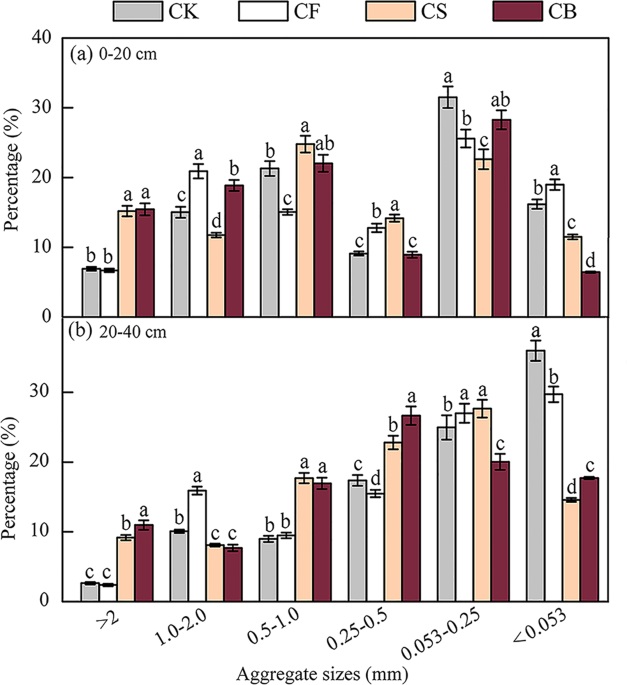
Soil water-stable aggregates were briefly separated using an agglomerate analyzer following the wet sieving method32,39,40. The sieve apertures used here were 2.00, 1.00, 0.50, 0.25, and 0.053 mm. Soil samples in the sieve were slowly submerged in sterilized water for 5 min. The analyzer was then vertically shaken for 5 min at 50 times/min with the column kept in water. The fractions remaining on each sieve and aggregates of diameter <0.053 mm settled in the sieve barrel were respectively collected at the end of sieving. The parameters of R0.25, MWD, GMD, and D were calculated as follows41,42:
$${{rm{R}}}_{0.25}=frac{{{rm{W}}}_{{rm{r}} > 0.25}}{{{rm{W}}}_{0}}$$
(1)
$${rm{MWD}}=mathop{sum }limits_{{rm{i}}=1}^{{rm{n}}}{{rm{X}}}_{{rm{i}}}{{rm{W}}}_{{rm{i}}}$$
(2)
$${rm{GMD}}=exp (frac{{sum }_{{rm{i}}=1}^{{rm{n}}}{{rm{W}}}_{{rm{i}}}{{rm{lnX}}}_{{rm{i}}}}{{sum }_{{rm{i}}=1}^{{rm{n}}}{{rm{W}}}_{{rm{i}}}})$$
(3)
$$(3-{rm{D}})mathrm{lg}left(frac{{{rm{X}}}_{{rm{i}}}}{{{rm{X}}}_{{rm{max }}}}right)=,{rm{lg}},frac{[{{rm{W}}}_{({rm{delta }}le {{rm{X}}}_{{rm{i}}})}]}{{{rm{W}}}_{0}}$$
(4)
where, R0.25 is the proportion of aggregates with a diameter of >0.25 mm; Xi refers to the mean diameter of i-size aggregates; Wi is the dry weight of the i-size fraction collected relative to the total soil used; Xmax, W(δ≤Xi), and W0 refer to the maximum diameter of all the aggregates tested, weight of aggregates with the particle-size <Xi, and total weight of each size fraction, respectively. Therefore, D could be obtained by regression analysis.
Soil water content was measured gravimetrically and expressed as a percentage of soil water to dry soil with constant weight. Soil pH was measured (soil:water = 1:2.5) using a precision pH meter (METTLER TOLEDO, Shanghai, China). SOC was determined using an elemental analyzer (ELEMENTAR, Langenselbold, Germany) after complete removal of the inorganic carbon in soil by 1 M HCl43. NH4+–N and NO3−–N were assayed using Nessler’s reagent and the phenol disulfonic acid colorimetric methods, respectively. AP was measured using the ammonium molybdate ascorbic method after extraction with 0.5 M NaHCO37. The CEC was determined using 1 M NH4OAc at pH 7.0 according to the protocol of the Analytical Methods of Soil Agricultural Chemistry44.
Soil DNA extraction and high-throughput sequencing
The total genomic DNA was extracted from 0.5 g of soil aggregates using the MOBIO PowerSoil Soil DNA Isolation Kit and purified according to the manufacturer’s instructions. After quantification using a K5500 Micro-Spectrophotometer (KAIAO, Beijing, China), DNA was subjected to PCR amplification of the V3-V4 variable fragments of 16S rRNA with the primer set of 338 F (5′-ACTCCTACGGGAGGCAGCAG-3′) and 806 R (5′-GGACTACHVGGGTATCTAAT-3′)45. Sequencing was performed using Illumina MiSeq PE300 at Majorbio Bio-pharm Technology (Shanghai, China).
Quality filtering of the raw reads was performed to obtain high-quality clean reads according to Cutadapt (V1.9.1, http://cutadapt.readthedocs.io/en/stable/). Chimera sequences were identified and removed using the UCHIME algorithm (http://www.drive5.com/usearch/manual/uchime_algo.html), and clean reads were finally obtained. OTUs were clustered using UPARSE software (UPARSE v7.0.1001, http://drive5.com/uparse/) (97% similarity) and analyzed against the SILVA database. PCR product purification, library construction, and data processing and analysis were conducted as described previously37. Sequences (raw reads) have been deposited into the NCBI Sequence Read Archive (SRA) database (Accession Number: SRP148725).
Statistical analysis
All comparative analyses were based on the normalized OTU abundance. Comparative analysis of soil physicochemical parameters and bacterial community structure between treatments was performed using a one-way ANOVA with the LSD test; results with p < 0.05 were considered statistically significant (SPSS 19.0, SPSS Corp., Chicago, IL, USA). The Chao1, Shannon, Simpson, and Simpsoneven indices were calculated using QIIME V1.9.1. PLS-DA, community barplot analysis, PERMANOVA (Bray-Curtis distances with 9999 permutations), and db-RDA (Weighted Unifrac) plots were generated using R V2.15.3. Other figures were produced using ORIGIN 9.0 (Originlab Corporation, Northampton, MA, USA).
Statement
Field trials were designed to analyze the long-term effects of different fertilization approaches (blank control, conventional inorganic fertilization, straw returning, and straw-derived biochar amendment) on soil physicochemical properties, soil aggregation, crop production, and soil microenvironment, etc. Data about the soil aggregate size and physicochemical properties (Figs. 1, 2, and Table 2) were important for microbial community structure analysis. This manuscript quoted some relevant data of PeerJ (doi:10.7717/peerj.6171).
Source: Ecology - nature.com


